Workflow Type: Galaxy
Open
Frozen
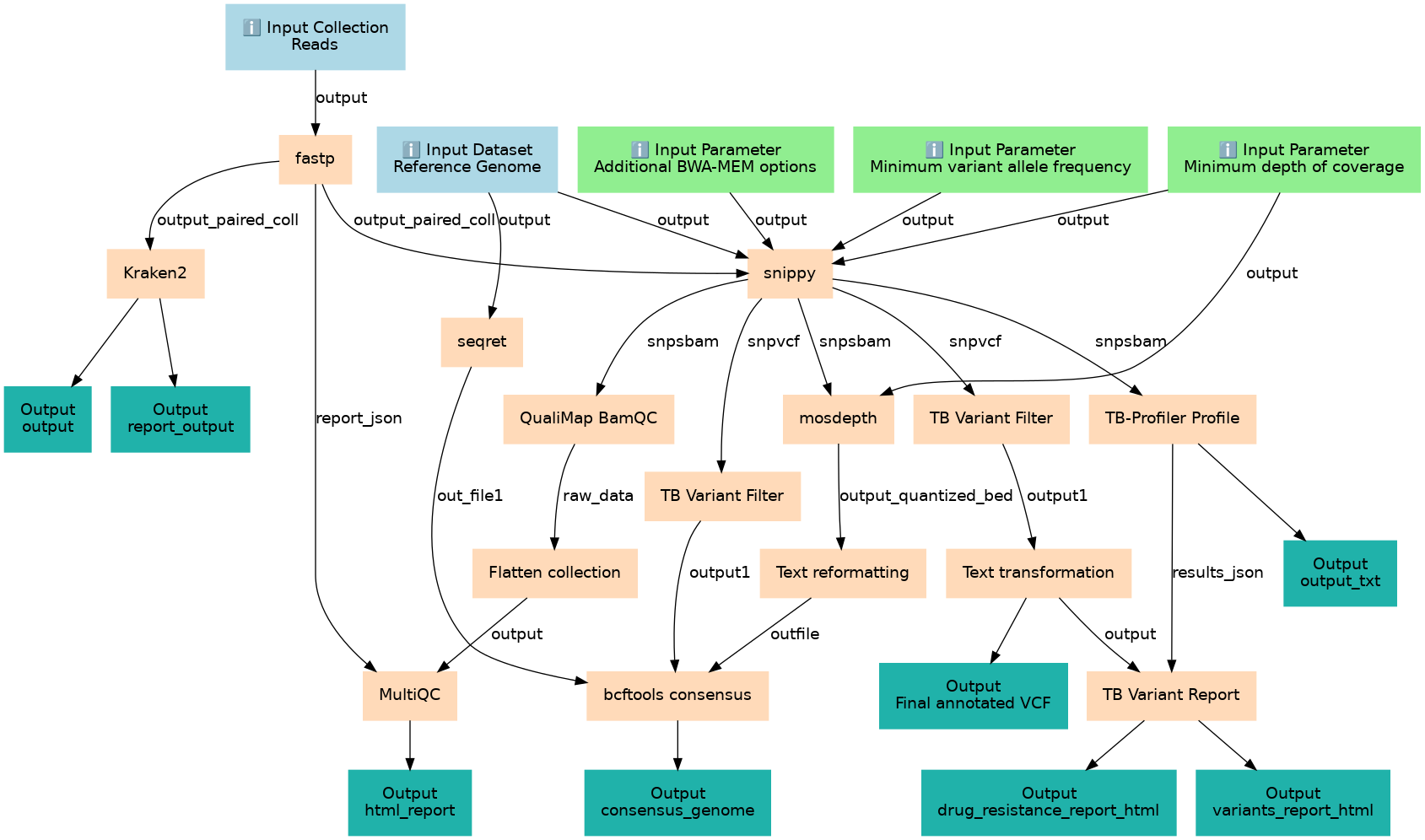
Predict variants and drug resistance from M. tuberculosis sequence samples (Illumina)
Associated Tutorial
This workflows is part of the tutorial TB Variant Analysis v1.0, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
Thanks to...
Tutorial Author(s): Peter van Heusden, Simon Gladman, Thoba Lose
Workflow Author(s): Peter van Heusden
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Additional BWA-MEM options | Additional BWA-MEM options | Increasing the minimum seed length (-k) parameter can help eliminate matches from contaminants. If used this should be set to 2/3rds of the read length, i.e. the string "-k 100" should be used for 150 base pair reads |
|
| Minimum depth of coverage | Minimum depth of coverage | Minimum depth of coverage for a position on the genome |
|
| Minimum variant allele frequency | Minimum variant allele frequency | Minimum proportion of reads that must support a variant for it to be called |
|
| Reads | Reads | List of paired Illumina reads (set format to fastqsanger or fastqsanger.gz as appropriate) |
|
| Reference Genome | Reference Genome | M. tuberculosis reference genome (must be in H37Rv coordinates. The M. tuberculosis inferred ancestral genome is recommended) |
|
Steps
| ID | Name | Description |
|---|---|---|
| 5 | fastp | toolshed.g2.bx.psu.edu/repos/iuc/fastp/fastp/0.23.4+galaxy0 |
| 6 | seqret | toolshed.g2.bx.psu.edu/repos/devteam/emboss_5/EMBOSS: seqret84/5.0.0 |
| 7 | snippy | toolshed.g2.bx.psu.edu/repos/iuc/snippy/snippy/4.6.0+galaxy0 |
| 8 | Kraken2 | toolshed.g2.bx.psu.edu/repos/iuc/kraken2/kraken2/2.1.1+galaxy1 |
| 9 | QualiMap BamQC | toolshed.g2.bx.psu.edu/repos/iuc/qualimap_bamqc/qualimap_bamqc/2.2.2c+galaxy1 |
| 10 | mosdepth | toolshed.g2.bx.psu.edu/repos/iuc/mosdepth/mosdepth/0.3.8+galaxy0 |
| 11 | TB Variant Filter | toolshed.g2.bx.psu.edu/repos/iuc/tb_variant_filter/tb_variant_filter/0.4.0+galaxy0 |
| 12 | TB-Profiler Profile | toolshed.g2.bx.psu.edu/repos/iuc/tbprofiler/tb_profiler_profile/6.2.1+galaxy0 |
| 13 | TB Variant Filter | toolshed.g2.bx.psu.edu/repos/iuc/tb_variant_filter/tb_variant_filter/0.4.0+galaxy0 |
| 14 | Flatten collection | __FLATTEN__ |
| 15 | Text reformatting | bcftools consensus requires a region file of low coverage regions in 1 based coordinates, mosdepth produces one in BED format with LOW and PASS for low coverage and non-low coverage regions. This converts from the one format to the other. toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_awk_tool/9.3+galaxy1 |
| 16 | Text transformation | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_sed_tool/9.3+galaxy1 |
| 17 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.11+galaxy1 |
| 18 | bcftools consensus | Compute consensus genome with single nucleotide variants inserted, suitable for use in building a phylogeny toolshed.g2.bx.psu.edu/repos/iuc/bcftools_consensus/bcftools_consensus/1.15.1+galaxy3 |
| 19 | TB Variant Report | toolshed.g2.bx.psu.edu/repos/iuc/tbvcfreport/tbvcfreport/1.0.0+galaxy0 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| report_output | report_output | n/a |
|
| output | output | n/a |
|
| output_txt | output_txt | n/a |
|
| Final annotated VCF | Final annotated VCF | n/a |
|
| html_report | html_report | n/a |
|
| consensus_genome | consensus_genome | n/a |
|
| drug_resistance_report_html | drug_resistance_report_html | n/a |
|
| variants_report_html | variants_report_html | n/a |
|
Version History
7.0 (latest) Created 16th Jul 2024 at 14:09 by Helena Rasche
Added/updated 4 files
Open
 master
masterd6f4ed5
14.0 (earliest) Created 25th Jun 2024 at 10:54 by Helena Rasche
Added/updated 4 files
Frozen
 14.0
14.0384a6af
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Activity
Views: 1340 Downloads: 551 Runs: 0
Created: 25th Jun 2024 at 10:54
Last updated: 25th Jun 2024 at 10:54
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992