Workflow Type: Galaxy
Open
Frozen
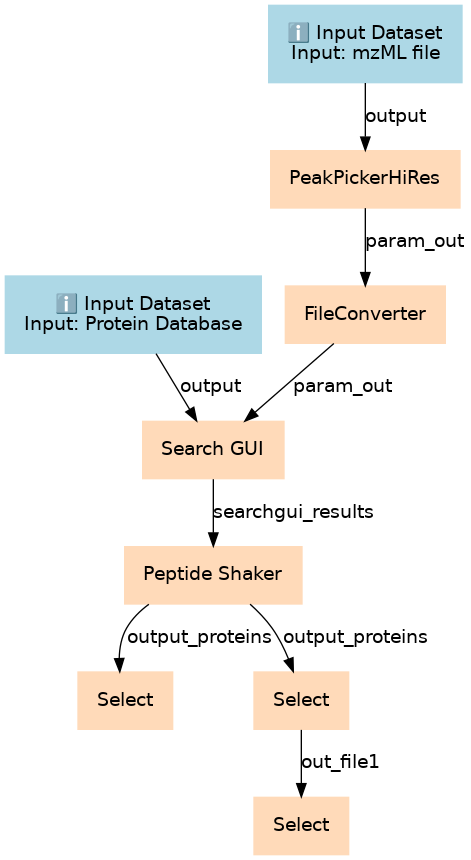
Peptide and Protein ID using SearchGUI and PeptideShaker
Associated Tutorial
This workflows is part of the tutorial Peptide And Protein ID Tutorial, available in the GTN
Thanks to...
Tutorial Author(s): Florian Christoph Sigloch, Björn Grüning
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Input: Protein Database | Input: Protein Database | n/a |
|
| Input: mzML file | Input: mzML file | n/a |
|
Steps
| ID | Name | Description |
|---|---|---|
| 2 | PeakPickerHiRes | PeakPickerHiRes |
| 3 | FileConverter | FileConverter |
| 4 | Search GUI | toolshed.g2.bx.psu.edu/repos/galaxyp/peptideshaker/search_gui/2.9.0 |
| 5 | Peptide Shaker | toolshed.g2.bx.psu.edu/repos/galaxyp/peptideshaker/peptide_shaker/1.11.0 |
| 6 | Select | Output: all identified contaminants. Grep1 |
| 7 | Select | Output: all identified proteins without common contaminants. CAVE: some proteins may be both! Grep1 |
| 8 | Select | Output: only those non-contaminant proteins not evaluated to be "Doubtful". Grep1 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| _anonymous_output_3 | _anonymous_output_3 | n/a |
|
| _anonymous_output_4 | _anonymous_output_4 | n/a |
|
| _anonymous_output_5 | _anonymous_output_5 | n/a |
|
| _anonymous_output_6 | _anonymous_output_6 | n/a |
|
| _anonymous_output_7 | _anonymous_output_7 | n/a |
|
| _anonymous_output_8 | _anonymous_output_8 | n/a |
|
| _anonymous_output_9 | _anonymous_output_9 | n/a |
|
| _anonymous_output_10 | _anonymous_output_10 | n/a |
|
Version History
4.0 (latest) Created 16th Jul 2024 at 14:07 by Helena Rasche
Added/updated 4 files
Open
 master
master4e0e0ad
5.0 (earliest) Created 25th Jun 2024 at 11:06 by Helena Rasche
Added/updated 4 files
Frozen
 5.0
5.0d9f70d5
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Activity
Views: 1451 Downloads: 555 Runs: 0
Created: 25th Jun 2024 at 11:06
Last updated: 25th Jun 2024 at 11:06
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992