Workflow Type: Galaxy
Open
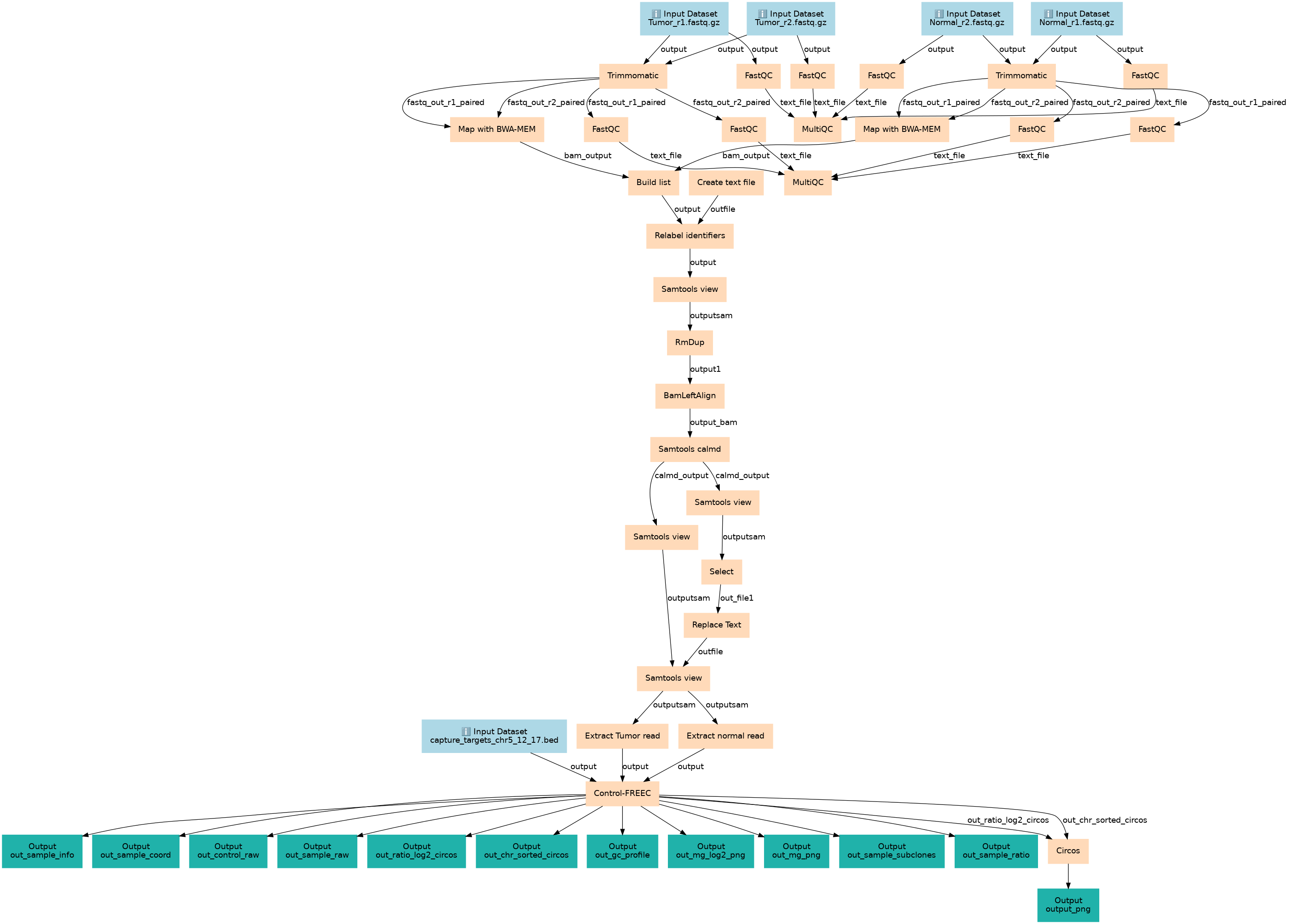
This workflow is created as part of a tutorial listed on GTN. The workflow shows the steps in human copy number variance detection using the Contrl_FREEC tool.
Associated Tutorial
This workflows is part of the tutorial Somatic-Variant-Discovery-from-WES-Data-Using-Control-FREEC, available in the GTN
Features
- Includes Galaxy Workflow Tests
Thanks to...
Tutorial Author(s): Khaled Jum'ah, Katarzyna Kamieniecka, Wolfgang Maier, David Salgado, Krzysztof Poterlowicz
Workflow Author(s): khaled Jumah, Katarzyna Kamieniecka, Wolfgang Maier, Krzysztof Poterlowicz, poterlowicz-lab
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Normal_r1.fastq.gz | Normal_r1.fastq.gz | n/a |
|
| Normal_r2.fastq.gz | Normal_r2.fastq.gz | n/a |
|
| Tumor_r1.fastq.gz | Tumor_r1.fastq.gz | n/a |
|
| Tumor_r2.fastq.gz | Tumor_r2.fastq.gz | n/a |
|
| capture_targets_chr5_12_17.bed | capture_targets_chr5_12_17.bed | n/a |
|
Steps
| ID | Name | Description |
|---|---|---|
| 3 | Create text file | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_text_file_with_recurring_lines/1.1.0 |
| 6 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 7 | Trimmomatic | toolshed.g2.bx.psu.edu/repos/pjbriggs/trimmomatic/trimmomatic/0.39+galaxy0 |
| 8 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 9 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 10 | Trimmomatic | toolshed.g2.bx.psu.edu/repos/pjbriggs/trimmomatic/trimmomatic/0.39+galaxy0 |
| 11 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 12 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 13 | Map with BWA-MEM | toolshed.g2.bx.psu.edu/repos/devteam/bwa/bwa_mem/0.7.17.2 |
| 14 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 15 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 16 | Map with BWA-MEM | toolshed.g2.bx.psu.edu/repos/devteam/bwa/bwa_mem/0.7.17.2 |
| 17 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.74+galaxy0 |
| 18 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.11+galaxy1 |
| 19 | Build list | __BUILD_LIST__ |
| 20 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.11+galaxy1 |
| 21 | Relabel identifiers | __RELABEL_FROM_FILE__ |
| 22 | Samtools view | toolshed.g2.bx.psu.edu/repos/iuc/samtools_view/samtools_view/1.15.1+galaxy0 |
| 23 | RmDup | toolshed.g2.bx.psu.edu/repos/devteam/samtools_rmdup/samtools_rmdup/2.0.1 |
| 24 | BamLeftAlign | toolshed.g2.bx.psu.edu/repos/devteam/freebayes/bamleftalign/1.3.6 |
| 25 | Samtools calmd | toolshed.g2.bx.psu.edu/repos/devteam/samtools_calmd/samtools_calmd/2.0.3 |
| 26 | Samtools view | toolshed.g2.bx.psu.edu/repos/iuc/samtools_view/samtools_view/1.15.1+galaxy0 |
| 27 | Samtools view | toolshed.g2.bx.psu.edu/repos/iuc/samtools_view/samtools_view/1.15.1+galaxy0 |
| 28 | Select | Grep1 |
| 29 | Replace Text | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_replace_in_line/1.1.2 |
| 30 | Samtools view | toolshed.g2.bx.psu.edu/repos/iuc/samtools_view/samtools_view/1.15.1+galaxy0 |
| 31 | Extract Tumor read | __EXTRACT_DATASET__ |
| 32 | Extract normal read | __EXTRACT_DATASET__ |
| 33 | Control-FREEC | toolshed.g2.bx.psu.edu/repos/iuc/control_freec/control_freec/11.6+galaxy2 |
| 34 | Circos | toolshed.g2.bx.psu.edu/repos/iuc/circos/circos/0.69.8+galaxy9 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| out_sample_info | out_sample_info | n/a |
|
| out_sample_coord | out_sample_coord | n/a |
|
| out_control_raw | out_control_raw | n/a |
|
| out_sample_raw | out_sample_raw | n/a |
|
| out_ratio_log2_circos | out_ratio_log2_circos | n/a |
|
| out_chr_sorted_circos | out_chr_sorted_circos | n/a |
|
| out_gc_profile | out_gc_profile | n/a |
|
| out_mg_log2_png | out_mg_log2_png | n/a |
|
| out_mg_png | out_mg_png | n/a |
|
| out_sample_subclones | out_sample_subclones | n/a |
|
| out_sample_ratio | out_sample_ratio | n/a |
|
| output_png | output_png | n/a |
|
Version History
1.0 (earliest) Created 25th Jun 2024 at 10:59 by Helena Rasche
Added/updated 4 files
Open
 master
master24fe8ae
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Activity
Views: 1088 Downloads: 312 Runs: 0
Created: 25th Jun 2024 at 10:59
Last updated: 25th Jun 2024 at 10:59
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992