Workflow Type: Galaxy
Open
Frozen
Protein FASTA Database Handling
Associated Tutorial
This workflows is part of the tutorial Proteomics: database handling, available in the GTN
Thanks to...
Tutorial Author(s): Florian Christoph Sigloch, Björn Grüning
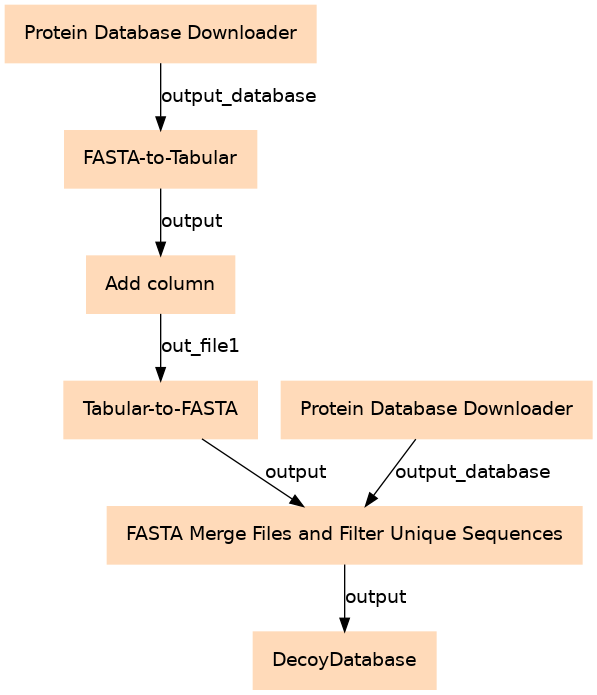
Steps
| ID | Name | Description |
|---|---|---|
| 0 | Protein Database Downloader | toolshed.g2.bx.psu.edu/repos/galaxyp/dbbuilder/dbbuilder/0.3.1 |
| 1 | Protein Database Downloader | toolshed.g2.bx.psu.edu/repos/galaxyp/dbbuilder/dbbuilder/0.3.1 |
| 2 | FASTA-to-Tabular | toolshed.g2.bx.psu.edu/repos/devteam/fasta_to_tabular/fasta2tab/1.1.1 |
| 3 | Add column | addValue |
| 4 | Tabular-to-FASTA | toolshed.g2.bx.psu.edu/repos/devteam/tabular_to_fasta/tab2fasta/1.1.1 |
| 5 | FASTA Merge Files and Filter Unique Sequences | toolshed.g2.bx.psu.edu/repos/galaxyp/fasta_merge_files_and_filter_unique_sequences/fasta_merge_files_and_filter_unique_sequences/1.2.0 |
| 6 | DecoyDatabase | toolshed.g2.bx.psu.edu/repos/galaxyp/openms_decoydatabase/DecoyDatabase/2.6+galaxy0 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| _anonymous_output_1 | #main/_anonymous_output_1 | n/a |
|
| _anonymous_output_2 | #main/_anonymous_output_2 | n/a |
|
| _anonymous_output_3 | #main/_anonymous_output_3 | n/a |
|
| _anonymous_output_4 | #main/_anonymous_output_4 | n/a |
|
| _anonymous_output_5 | #main/_anonymous_output_5 | n/a |
|
| _anonymous_output_6 | #main/_anonymous_output_6 | n/a |
|
| _anonymous_output_7 | #main/_anonymous_output_7 | n/a |
|
Version History
5.0 (latest) Created 16th Jul 2024 at 14:07 by Helena Rasche
Added/updated 4 files
Open
 master
master16af6db
7.0 (earliest) Created 25th Jun 2024 at 11:09 by Helena Rasche
Added/updated 4 files
Frozen
 7.0
7.0248d62a
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Activity
Views: 1525 Downloads: 620 Runs: 0
Created: 25th Jun 2024 at 11:09
Last updated: 25th Jun 2024 at 11:09
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992