Workflow Type: Galaxy
Open
Frozen
Training workflow
Associated Tutorial
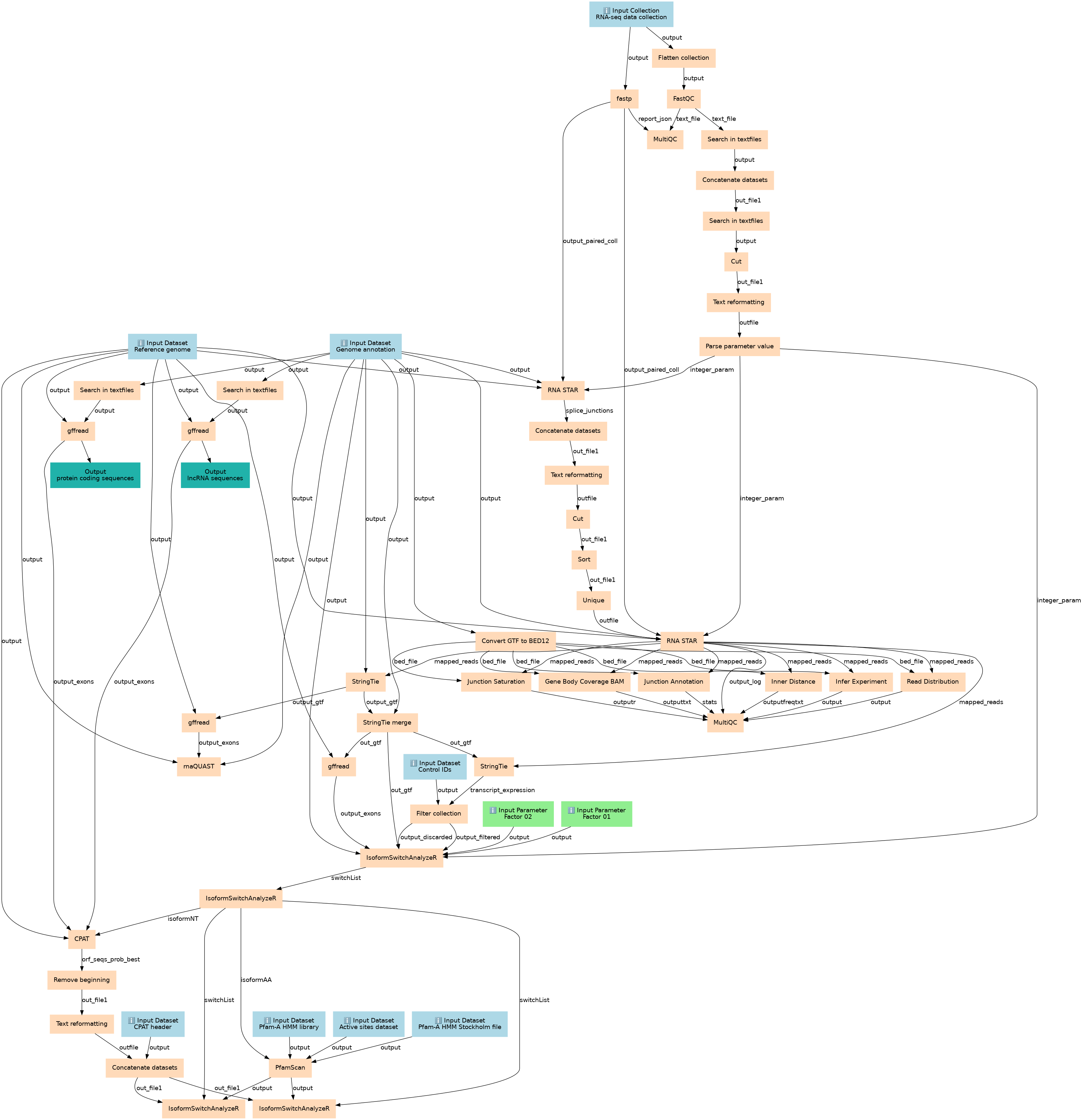
This workflows is part of the tutorial Genome-wide alternative splicing analysis: human, available in the GTN
Features
- Includes Galaxy Workflow Tests
Thanks to...
Tutorial Author(s): Cristóbal Gallardo, Lucille Delisle
Tutorial Contributor(s): Pavankumar Videm
Workflow Author(s): Cristóbal Gallardo Alba
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Active sites dataset | Active sites dataset | Active sites dataset. |
|
| CPAT header | CPAT header | Galaxy compatible CPAT header. |
|
| Control IDs | Control IDs | IDs of control sequences, corresponding to filenames. |
|
| Factor 01 | Factor 01 | First factor name. |
|
| Factor 02 | Factor 02 | Second factor name. |
|
| Genome annotation | Genome annotation | Reference genome annotation in GTF format. |
|
| Pfam-A HMM Stockholm file | Pfam-A HMM Stockholm file | Stockholm file. |
|
| Pfam-A HMM library | Pfam-A HMM library | HMM library. |
|
| RNA-seq data collection | RNA-seq data collection | Collection of paired-end RNA-seq datasets. |
|
| Reference genome | Reference genome | Genome reference in FASTA format. |
|
Steps
| ID | Name | Description |
|---|---|---|
| 10 | fastp | toolshed.g2.bx.psu.edu/repos/iuc/fastp/fastp/0.23.2+galaxy0 |
| 11 | Flatten collection | __FLATTEN__ |
| 12 | Search in textfiles | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_grep_tool/1.1.1 |
| 13 | Search in textfiles | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_grep_tool/1.1.1 |
| 14 | Convert GTF to BED12 | toolshed.g2.bx.psu.edu/repos/iuc/gtftobed12/gtftobed12/357 |
| 15 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.73+galaxy0 |
| 16 | gffread | toolshed.g2.bx.psu.edu/repos/devteam/gffread/gffread/2.2.1.3+galaxy0 |
| 17 | gffread | toolshed.g2.bx.psu.edu/repos/devteam/gffread/gffread/2.2.1.3+galaxy0 |
| 18 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.11+galaxy1 |
| 19 | Search in textfiles | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_grep_tool/1.1.1 |
| 20 | Concatenate datasets | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_cat/0.1.1 |
| 21 | Search in textfiles | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_grep_tool/1.1.1 |
| 22 | Cut | Cut1 |
| 23 | Text reformatting | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_awk_tool/1.1.2 |
| 24 | Parse parameter value | param_value_from_file |
| 25 | RNA STAR | toolshed.g2.bx.psu.edu/repos/iuc/rgrnastar/rna_star/2.7.10b+galaxy3 |
| 26 | Concatenate datasets | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_cat/0.1.1 |
| 27 | Text reformatting | ($5 > 0 && $7 > 3 && $6==0) toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_awk_tool/1.1.2 |
| 28 | Cut | Cut1 |
| 29 | Sort | sort1 |
| 30 | Unique | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_sorted_uniq/1.1.0 |
| 31 | RNA STAR | toolshed.g2.bx.psu.edu/repos/iuc/rgrnastar/rna_star/2.7.10b+galaxy3 |
| 32 | StringTie | toolshed.g2.bx.psu.edu/repos/iuc/stringtie/stringtie/2.2.1+galaxy1 |
| 33 | Junction Annotation | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_junction_annotation/5.0.1+galaxy2 |
| 34 | Junction Saturation | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_junction_saturation/5.0.1+galaxy2 |
| 35 | Gene Body Coverage (BAM) | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_geneBody_coverage/5.0.1+galaxy2 |
| 36 | Inner Distance | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_inner_distance/5.0.1+galaxy2 |
| 37 | Infer Experiment | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_infer_experiment/5.0.1+galaxy2 |
| 38 | Read Distribution | toolshed.g2.bx.psu.edu/repos/nilesh/rseqc/rseqc_read_distribution/5.0.1+galaxy2 |
| 39 | gffread | toolshed.g2.bx.psu.edu/repos/devteam/gffread/gffread/2.2.1.3+galaxy0 |
| 40 | StringTie merge | toolshed.g2.bx.psu.edu/repos/iuc/stringtie/stringtie_merge/2.2.1+galaxy1 |
| 41 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.11+galaxy1 |
| 42 | rnaQUAST | toolshed.g2.bx.psu.edu/repos/iuc/rnaquast/rna_quast/2.2.3+galaxy0 |
| 43 | gffread | toolshed.g2.bx.psu.edu/repos/devteam/gffread/gffread/2.2.1.3+galaxy0 |
| 44 | StringTie | toolshed.g2.bx.psu.edu/repos/iuc/stringtie/stringtie/2.2.1+galaxy1 |
| 45 | Filter collection | __FILTER_FROM_FILE__ |
| 46 | IsoformSwitchAnalyzeR | toolshed.g2.bx.psu.edu/repos/iuc/isoformswitchanalyzer/isoformswitchanalyzer/1.20.0+galaxy5 |
| 47 | IsoformSwitchAnalyzeR | toolshed.g2.bx.psu.edu/repos/iuc/isoformswitchanalyzer/isoformswitchanalyzer/1.20.0+galaxy5 |
| 48 | CPAT | toolshed.g2.bx.psu.edu/repos/bgruening/cpat/cpat/3.0.4+galaxy0 |
| 49 | PfamScan | toolshed.g2.bx.psu.edu/repos/bgruening/pfamscan/pfamscan/1.6+galaxy0 |
| 50 | Remove beginning | Remove beginning1 |
| 51 | Text reformatting | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_awk_tool/1.1.2 |
| 52 | Concatenate datasets | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_cat/0.1.1 |
| 53 | IsoformSwitchAnalyzeR | toolshed.g2.bx.psu.edu/repos/iuc/isoformswitchanalyzer/isoformswitchanalyzer/1.20.0+galaxy5 |
| 54 | IsoformSwitchAnalyzeR | toolshed.g2.bx.psu.edu/repos/iuc/isoformswitchanalyzer/isoformswitchanalyzer/1.20.0+galaxy5 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| lncRNA sequences | lncRNA sequences | n/a |
|
| protein coding sequences | protein coding sequences | n/a |
|
Version History
3.0 (latest) Created 16th Sep 2024 at 14:07 by Helena Rasche
Added/updated 4 files
Open
 master
master1b3f813
14.0 (earliest) Created 25th Jun 2024 at 11:24 by Helena Rasche
Added/updated 4 files
Frozen
 14.0
14.0539ac6b
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Tools
Activity
Views: 1805 Downloads: 797 Runs: 0
Created: 25th Jun 2024 at 11:24
Last updated: 16th Sep 2024 at 14:07
 Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992