Workflows
What is a Workflow?Filters
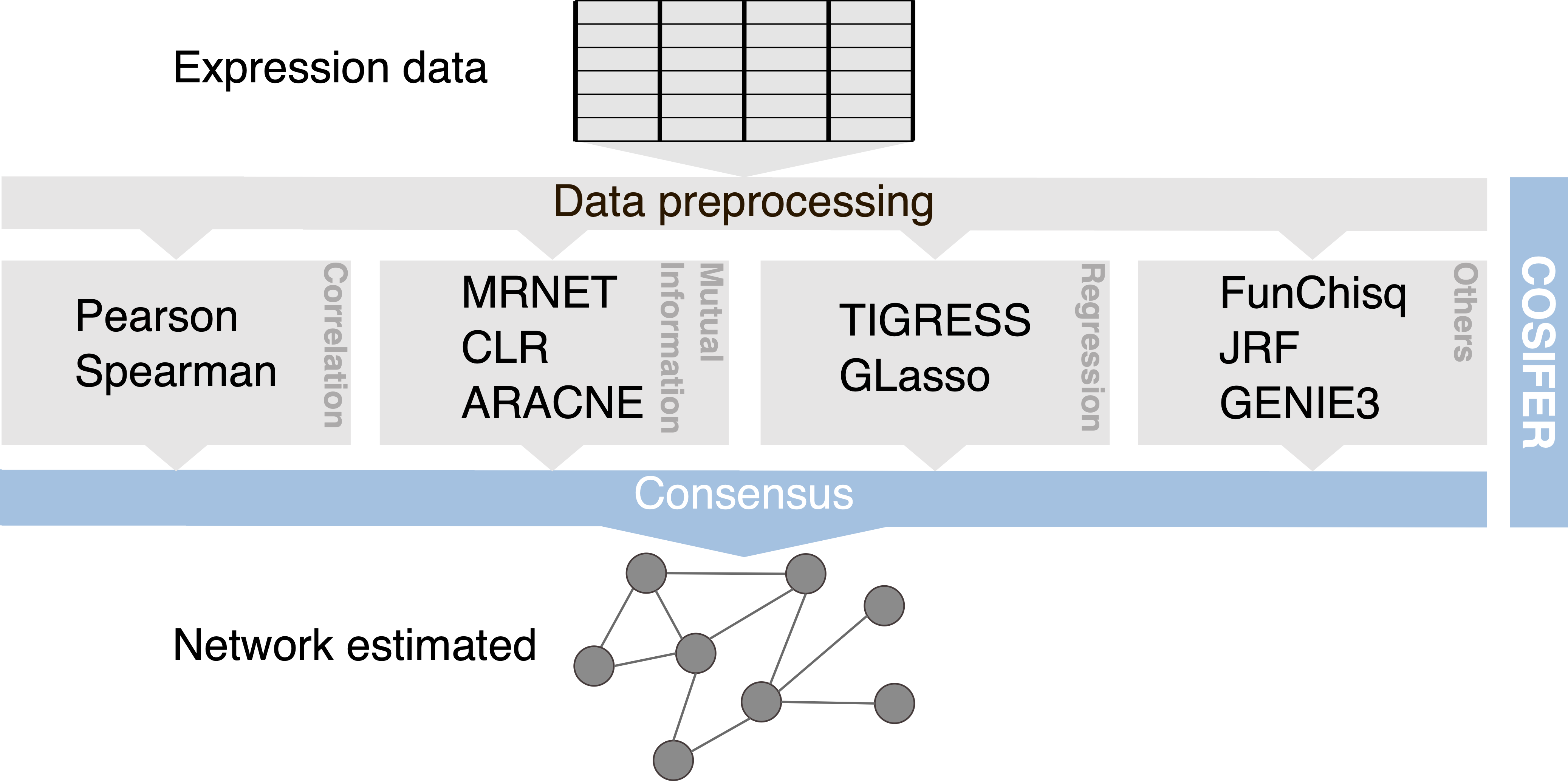
COnSensus Interaction Network InFErence Service
Inference framework for reconstructing networks using a consensus approach between multiple methods and data sources.
Reference
[Manica, Matteo, Charlotte, Bunne, Roland, Mathis, Joris, Cadow, Mehmet Eren, Ahsen, Gustavo A, Stolovitzky, and María Rodríguez, Martínez. "COSIFER: a python package for the consensus inference of molecular ...
COnSensus Interaction Network InFErence Service
Inference framework for reconstructing networks using a consensus approach between multiple methods and data sources.

Reference
[Manica, Matteo, Charlotte, Bunne, Roland, Mathis, Joris, Cadow, Mehmet Eren, Ahsen, Gustavo A, Stolovitzky, and María Rodríguez, Martínez. "COSIFER: a python package for the consensus inference of molecular ...
Type: Common Workflow Language
Creators: José Mª Fernández, Laura Rodriguez-Navas
Submitter: Laura Rodriguez-Navas
COVID-19: variation analysis on ARTIC ONT data
This workflow for ONT-sequenced ARTIC data is modeled after the alignment/variant-calling steps of the ARTIC pipeline. It performs, essentially, the same steps as that pipeline’s minion command, i.e. read mapping with minimap2 and variant calling with medaka. Like the Illumina ARTIC workflow it uses ivar for primer trimming. Since ONT-sequenced reads have a much ...
COVID-19: variation analysis on WGS PE data
This workflows performs single end read mapping with bowtie2 followed by sensitive variant calling across a wide range of AFs with lofreq and variant annotation with snpEff 4.5covid19.
COVID-19: variation analysis on WGS PE data
This workflows performs paired end read mapping with bwa-mem followed by sensitive variant calling across a wide range of AFs with lofreq and variant annotation with snpEff 4.5covid19.
COVID-19: variation analysis on ARTIC PE data
The workflow for Illumina-sequenced ARTIC data builds on the RNASeq workflow for paired-end data using the same steps for mapping and variant calling, but adds extra logic for trimming ARTIC primer sequences off reads with the ivar package. In addition, this workflow uses ivar also to identify amplicons affected by ARTIC primer-binding site mutations and excludes reads derived from such "tainted" amplicons ...
- Name of the application: Adapter removal hello world
- Name of the application: Mapping and bam sorting goodbye world
1. Name of the application: Adapter removal hello world 2. Name of the application: Mapping and bam sorting goodbye world
 Download
Download